Humans and other multicellular organisms are composed of a variety of organs, tissues, and cell types. Given that all cells share the same genome, why does such diversity arise? Why do cells arranged in three dimensions exhibit different functions depending on their location?
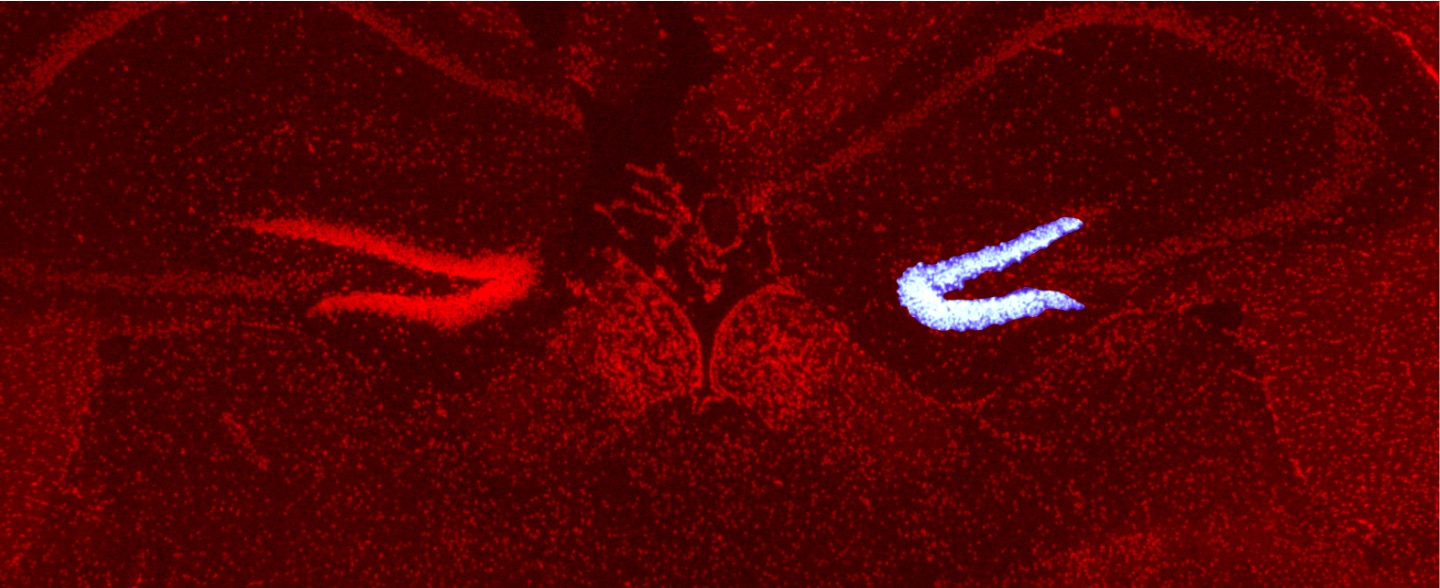
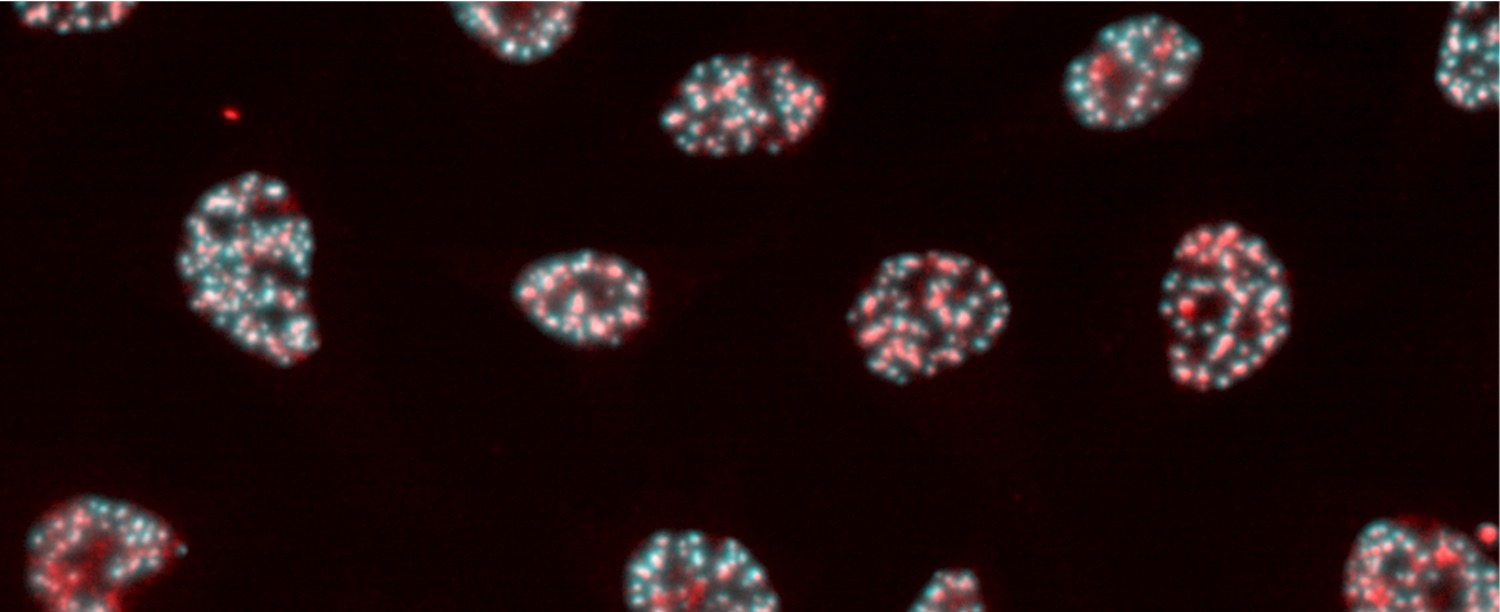
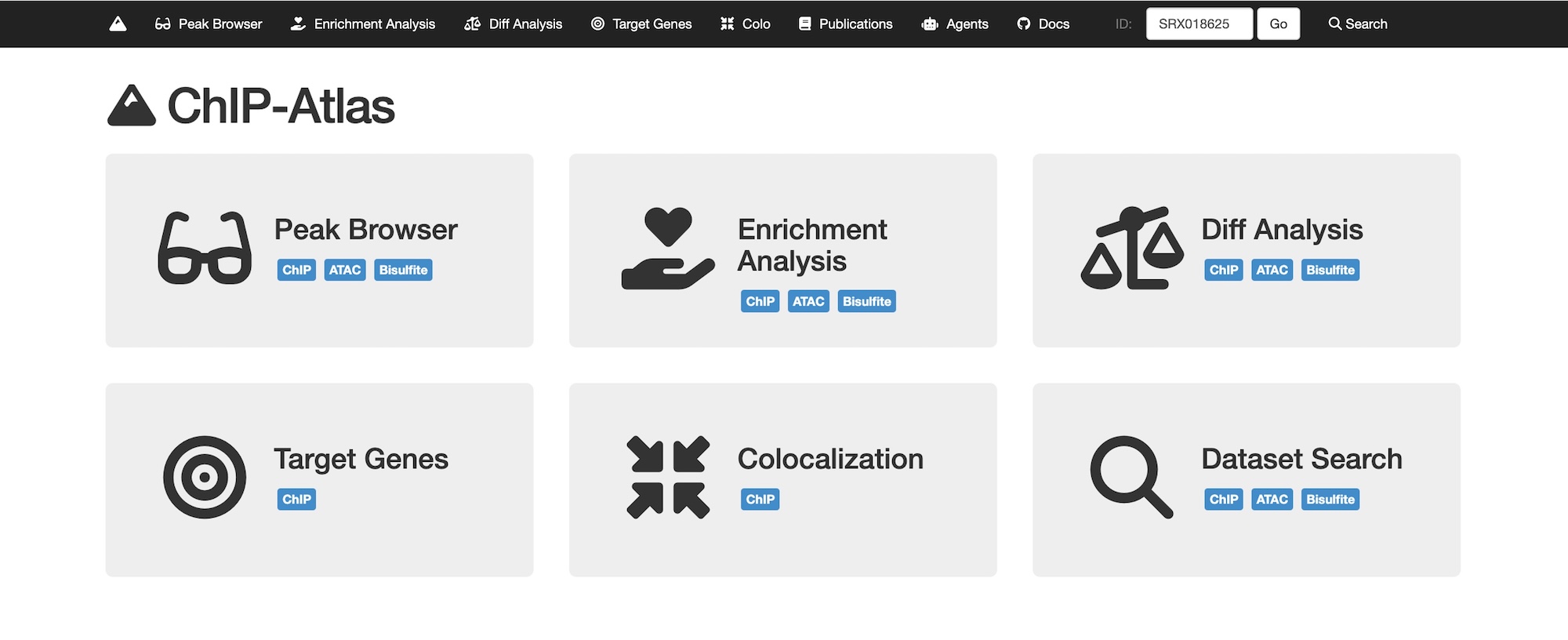
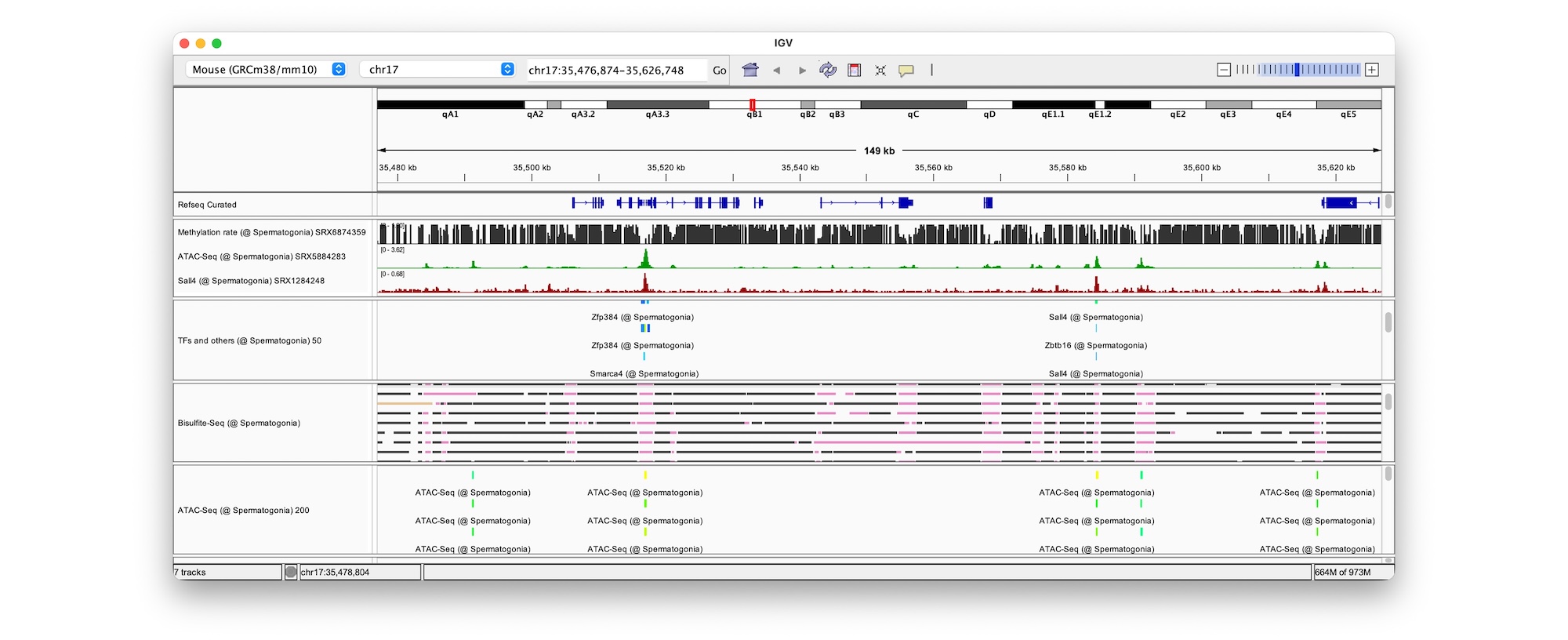
The key to solving this mystery lies in spatiotemporal gene expression. To address this, we developed PIC, a technology to profile transcriptomes in locally defined regions, and ChIP-Atlas, an integrative epigenome database for analyzing transcriptional regulation. By combining these approaches, we aim to elucidate disease onset mechanisms and drug actions.
Theme 2
What's New
- 2026-05-07A collaborative research paper with Dr. Atsuta (Kyushu Univ) has been published in Nature Communications. Zhaonan performed PIC RNA-seq on sternal precursor cells from emu and chicken embryos and factors determining species differences in sternal morphology. See the UV-irradiation scene here.
- 2026-05-01Dr. Omnia REDA has joined the Oki Lab as a postdoc researcher!
- 2026-04-30A paper by Zhaonan and Shinya describing ChIP-Atlas 2025 Update, has been published in Nucleic Acids Research! We have implemented a quality control framework for the experimental data and new analysis tools for RNA-seq upstream analysis. We thank our collaborators, Dr. Tazro Ohta (Chiba Univ) and Dr. Takeya Kasukawa (RIKEN).
- 2026-04-22English website is now live!