
Research
Decoding Spatiotemporal Gene Expression
At Okilab, our research focus is to elucidate the mechanisms of spatiotemporal gene expression in processes such as ontogenesis, cell differentiation, and disease onset. To achieve this, we are developing the necessary experimental techniques and data infrastructure. We have developed a spatial transcriptomics technology called PIC to comprehensively analyze genes that are expressed locally. In addition, we have developed ChIP-Atlas, an integrative epigenomics database that uses a data-driven approach to elucidate the mechanisms of gene expression regulation. By combining these approaches, we aim to elucidate disease onset mechanisms and drug actions from the perspective of spatiotemporal gene expression regulation.
Theme 1:Spatial transcriptomics using PIC
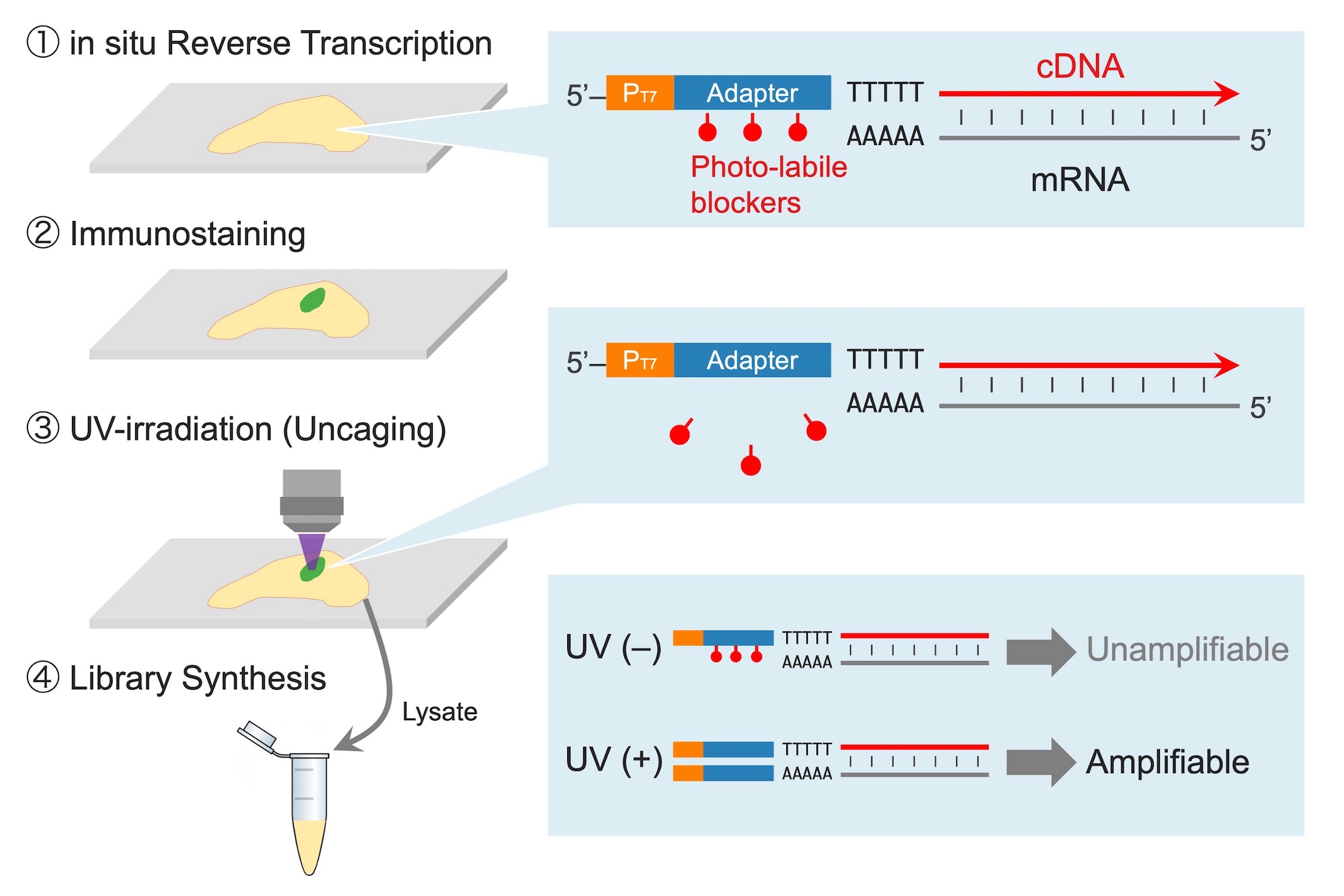
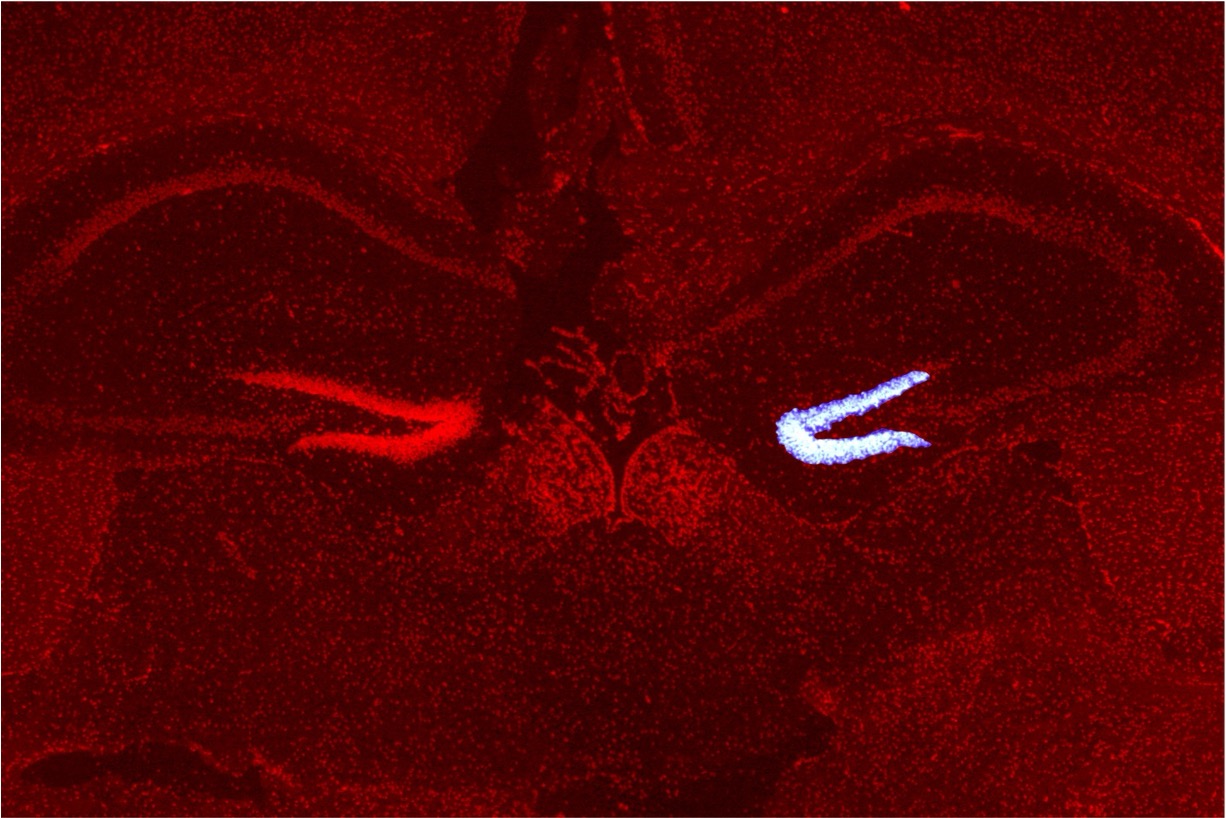
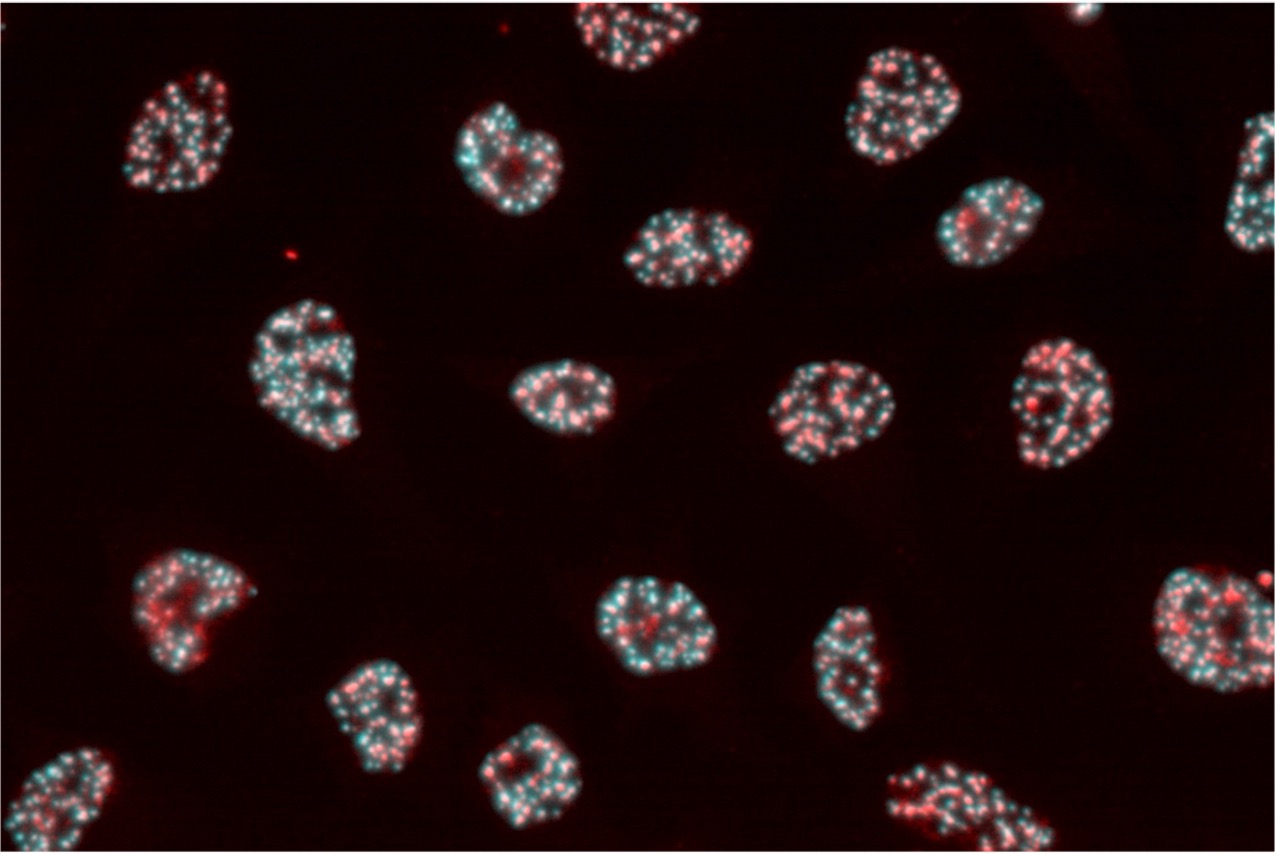
We have developed a technology called Photo-Isolation Chemistry (PIC) that can capture transcriptome within small cell populations and regions with world-leading spatial resolution and detection sensitivity (Honda et al., Nat Commun., 2021). This groundbreaking technology allows for the thorough detection of genes expressed in target cell populations or microtissues when exposed to UV. Using this method, we successfully detected genes that were specifically expressed in various regions of the brain by UV-irradiating those regions. Furthermore, we have achieved comprehensive gene detection in extremely small cell populations within mouse embryos and in submicron-scale structures within cell nuclei. We are currently developing PIC further to advance the technology for capturing epigenomic and proteomic information in microscopic regions besides transcriptomes.
Theme 2:Data mining using ChIP-Atlas
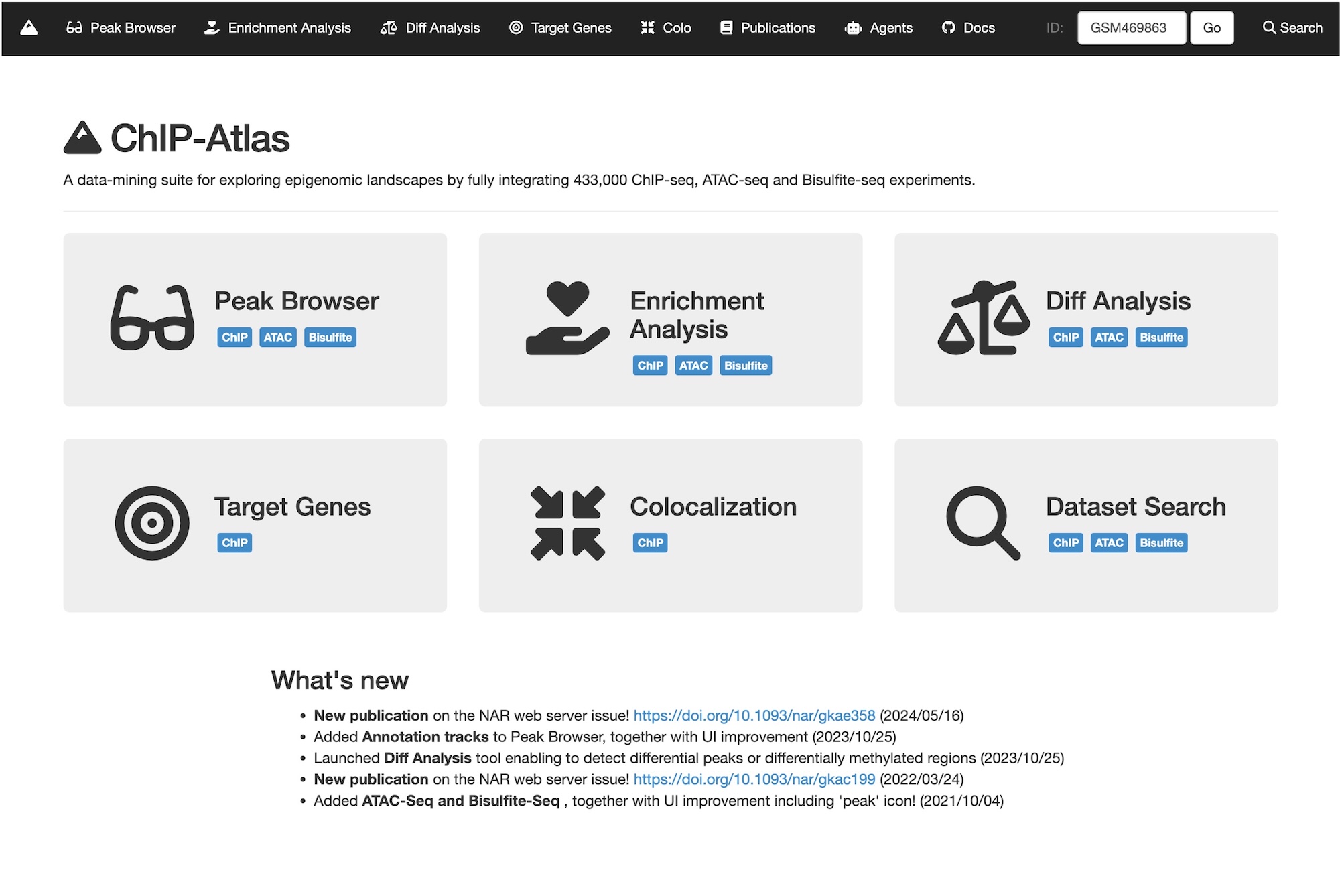
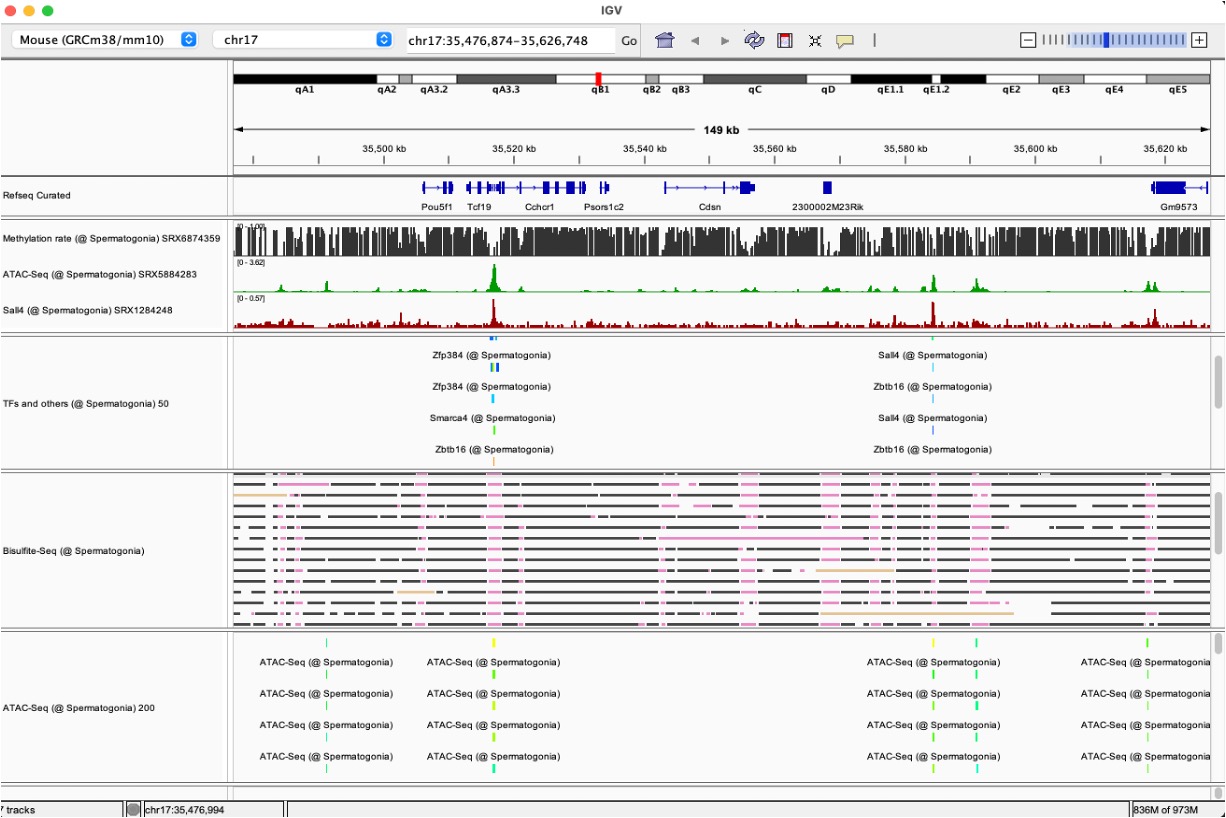
We are developing ChIP-Atlas, a database that integrates ChIP-seq data from around the world. Through comprehensive analyses of nearly 300,000 ChIP-seq experiments targeting modified histone marks and transcription factors, we identified over 20 billion protein–genome interactions. Recent updates have incorporated hundreds of thousands of ATAC-seq and Bisulfite-seq datasets, providing a comprehensive understanding of chromatin accessibility and DNA methylation status.
This vast dataset, spanning hundreds of terabytes, is publicly available through the ChIP-Atlas website, where it can be easily browsed and analyzed. Since its public release a decade ago, ChIP-Atlas has been widely used by researchers worldwide and cited in over 1,500 papers. Our goal is to use this resource to elucidate the epigenetic mechanisms that regulate spatiotemporal gene expression through a data-driven approach.
Theme 3:Integrative wet–dry applied research
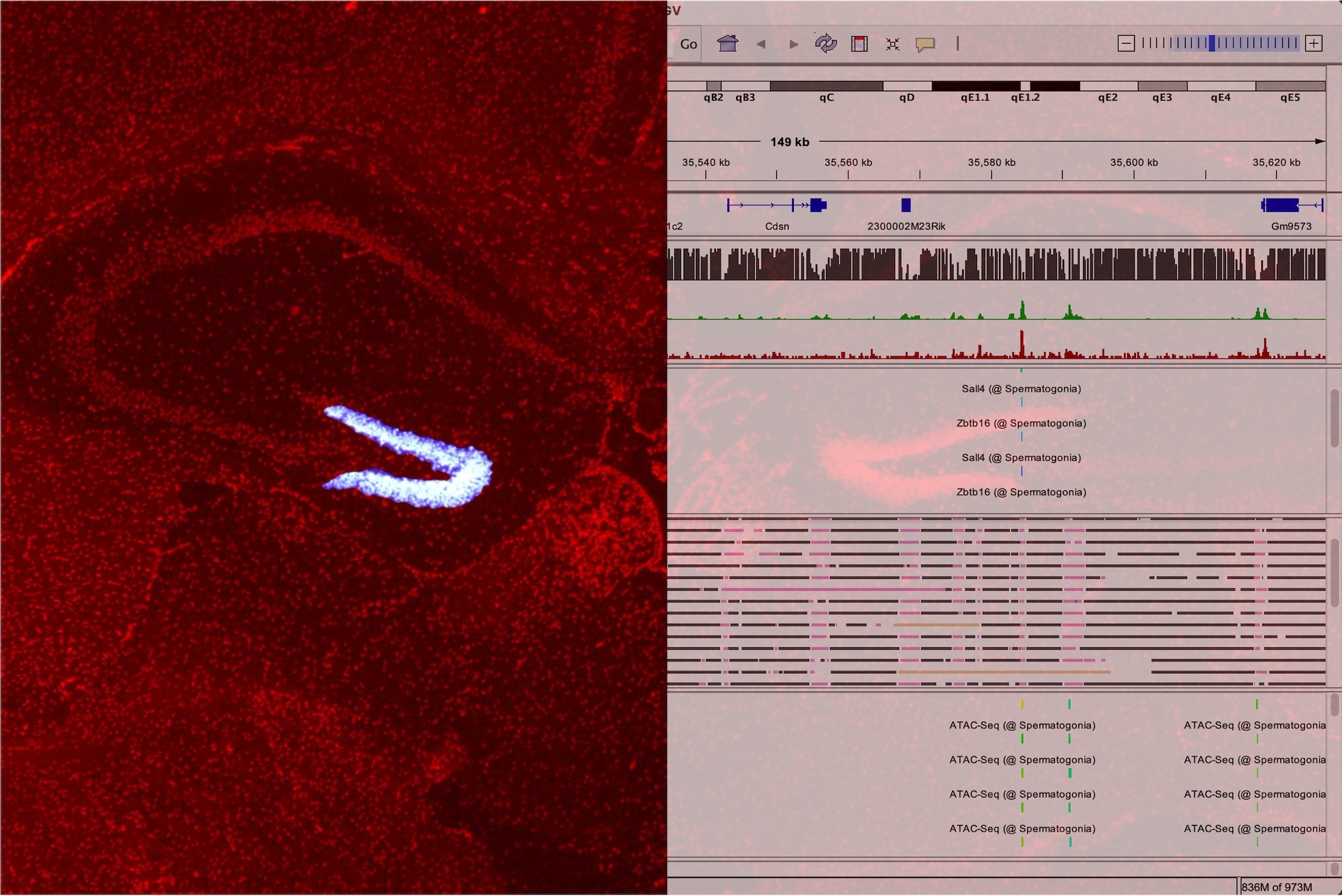
PIC can identify cell-type–specific gene expression in diverse organ or tissue. Furthermore, by analyzing these gene sets with ChIP-Atlas, we can identify the upstream TFs that integrally regulate them. This enables us to pinpoint the master regulatory factors that govern the differentiation of various cell types, raising hopes for applications in direct cell programming and regenerative medicine.
We are also analyzing gene sets whose expression fluctuates in response to drug administration. Using ChIP-Atlas, we can identify the pivotal regulatory factors of these drug-responsive genes. In other words, by understanding the TFs at the core of a drug's action modes, we can apply this knowledge to identify drug targets and predict new pharmacological effects. In addition, we are using ChIP-Atlas to determine which TFs bind to SNPs that cause hereditary diseases. This enables us to identify TFs involved in disease onset, which can inform the selection of effective drug candidates.